I-value: Supplementary Material
Citation: J.H. Collier, L.Allison, A.M. Lesk, M. Garcia de la Banda and A.S. Konagurthu. A new statistical framework to assess structural alignment quality using information compression. Bioinformatics (2014) 30 (17): i512-i518.
A new statistical framework to assess structural alignment quality using information compression
James H. Collier, Lloyd Allison, Arthur M. Lesk, Maria Garcia de la Banda, Arun S. Konagurthu
Raw score data of all alignment programs on the SCOP data described in Section 3 of the main text
Click on the links below to download raw score data for all scoring functions used. Each file contains score data for the respective alignment programs used on the 2500 structure pairs from SCOP. The first line in each file contains a column heading reference indicating which scores are in each column.
Table S1: Boxplots of scores/aligners on SCOP data in addition to Fig 1 of the main text.
Table displaying all alignment programs and scores including those in the main text.
| Aligner\Score | Dali Score | Dali Z-Score | TM-Score | GDT_TS | GSAS | LGA_S3 | MI | SAS | SI | Structal Score | I-value | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Dali | 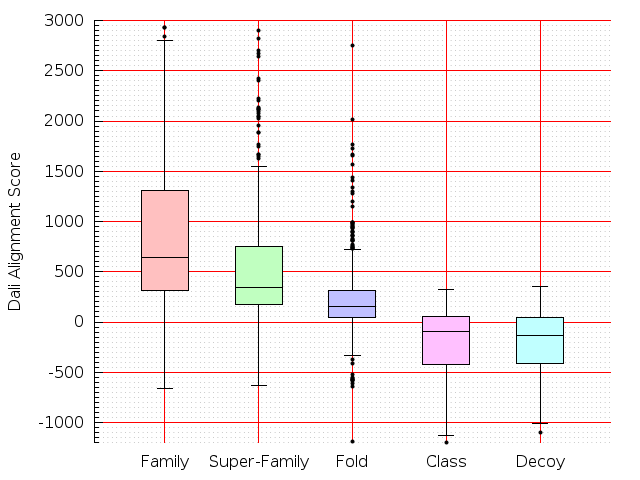 | 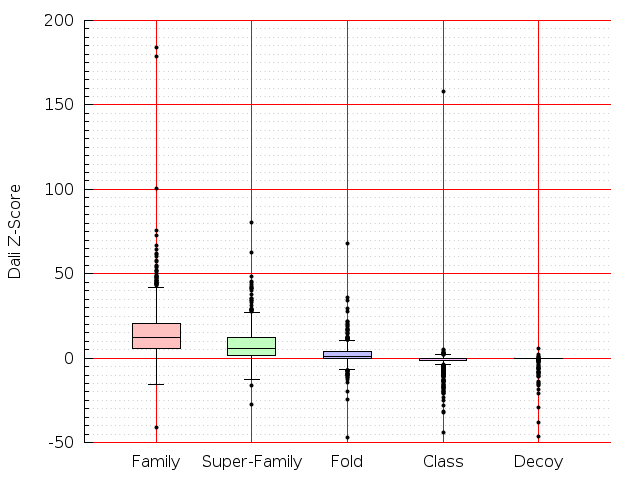 | 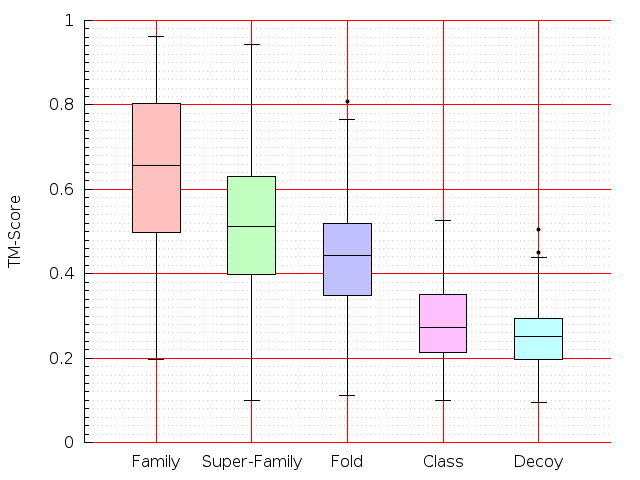 | 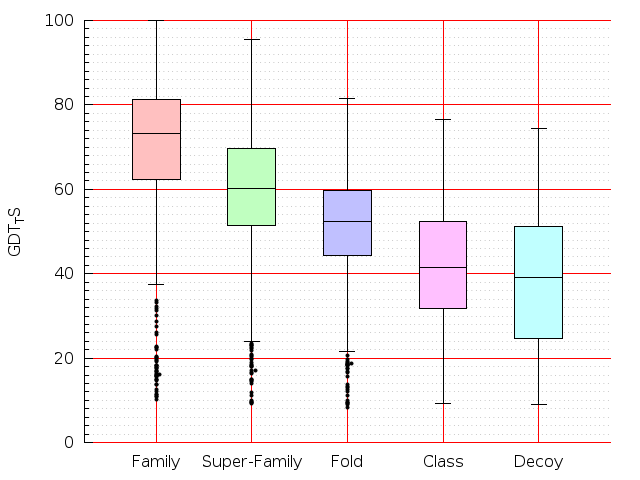 | 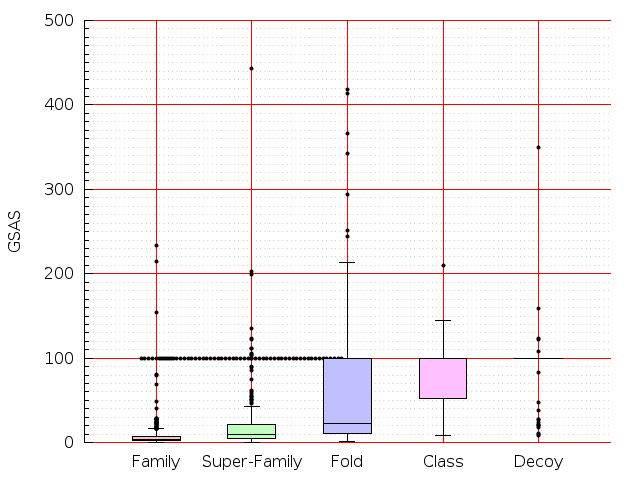 | 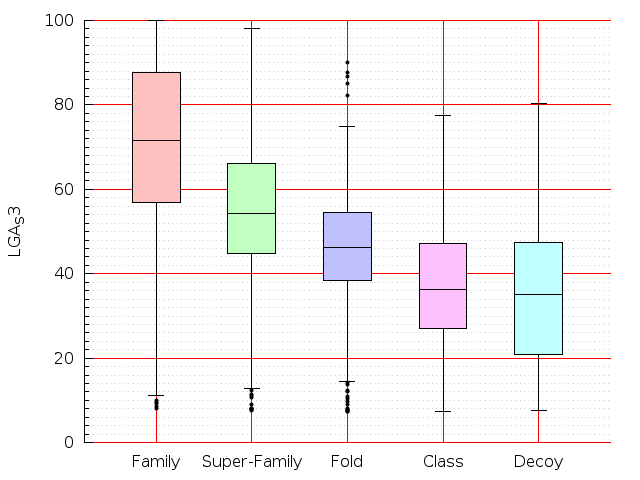 | 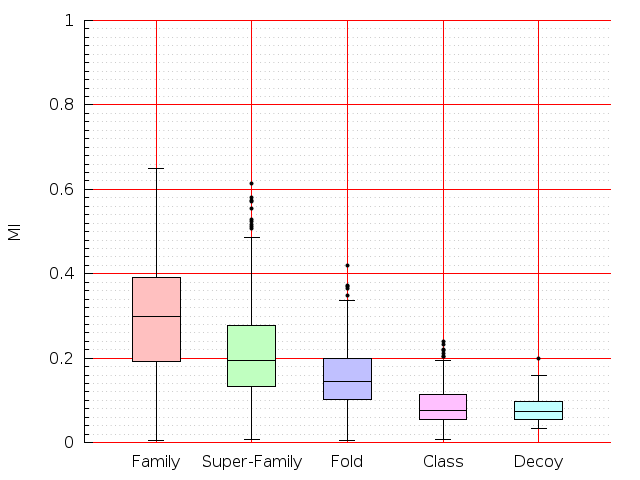 | 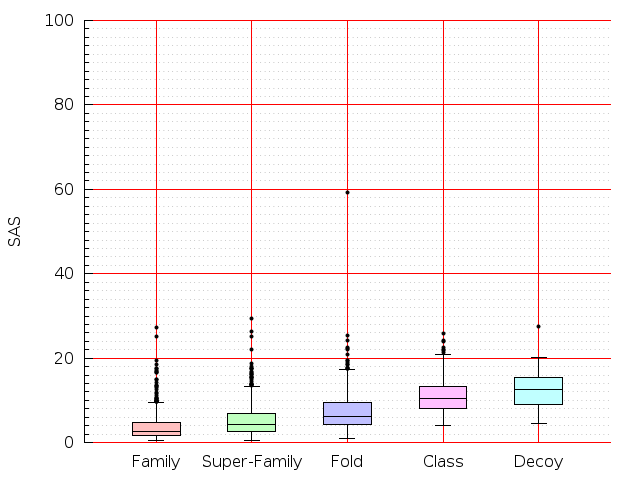 | 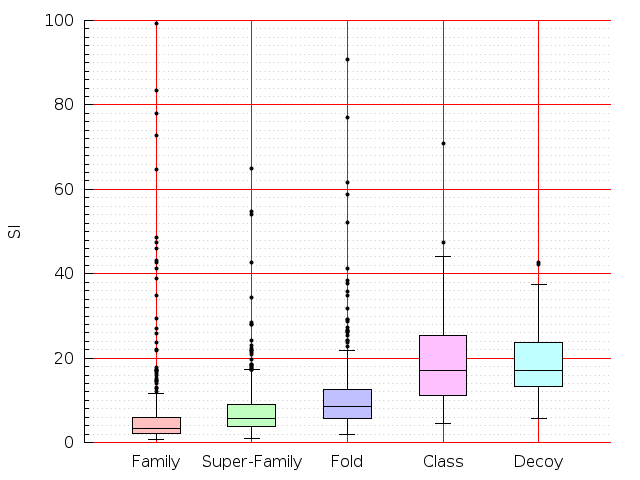 | 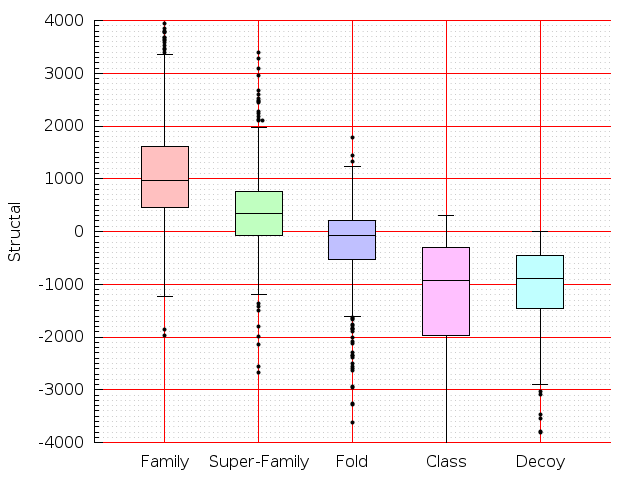 | 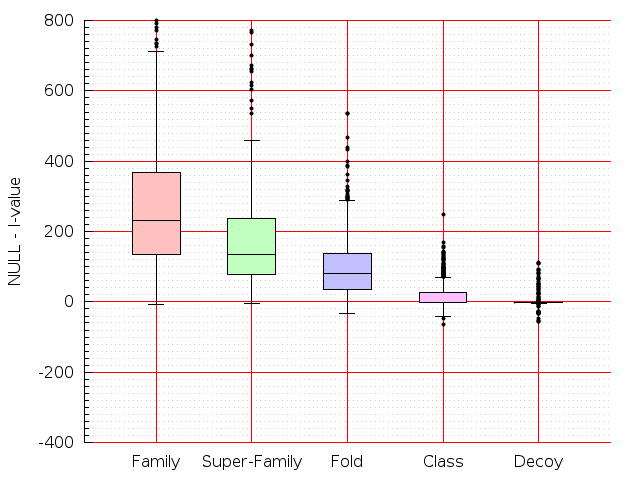 | Dali |
| TM-Align | 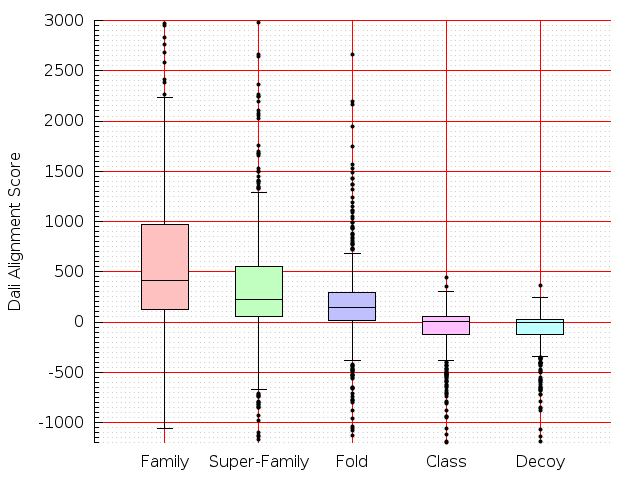 | 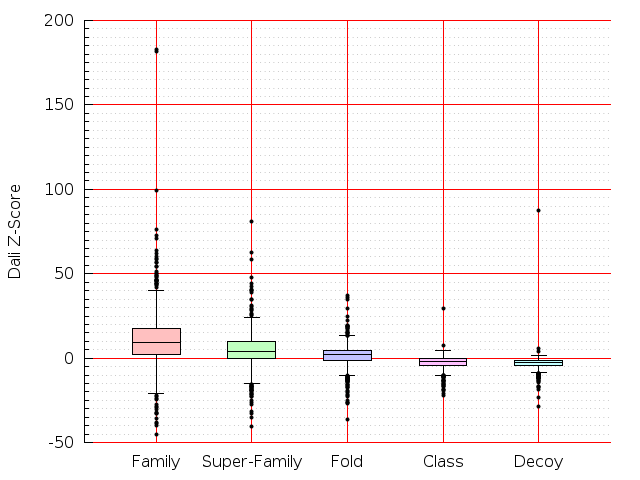 | 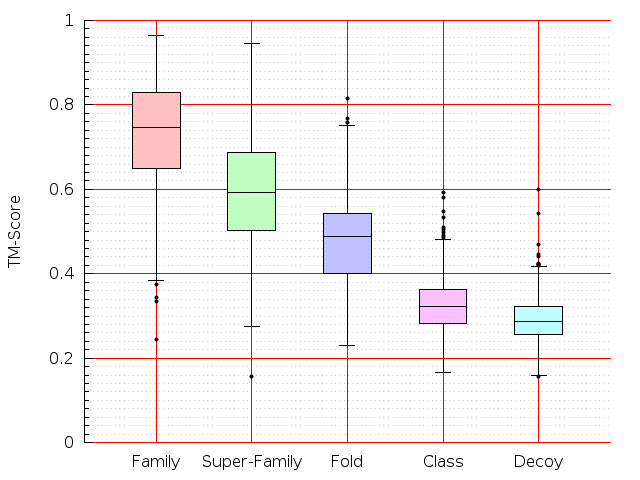 | 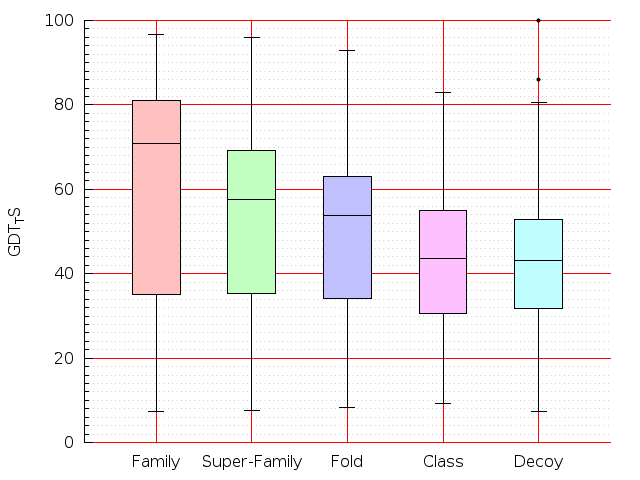 | 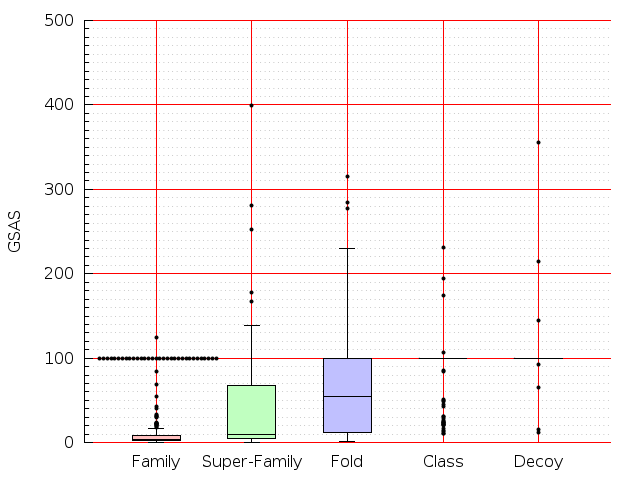 | 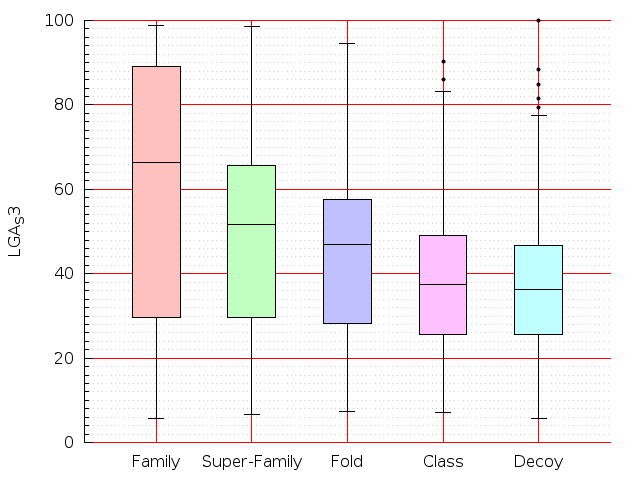 | 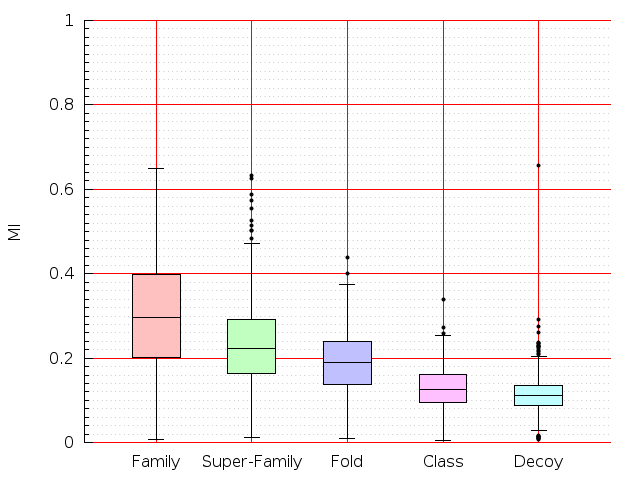 | 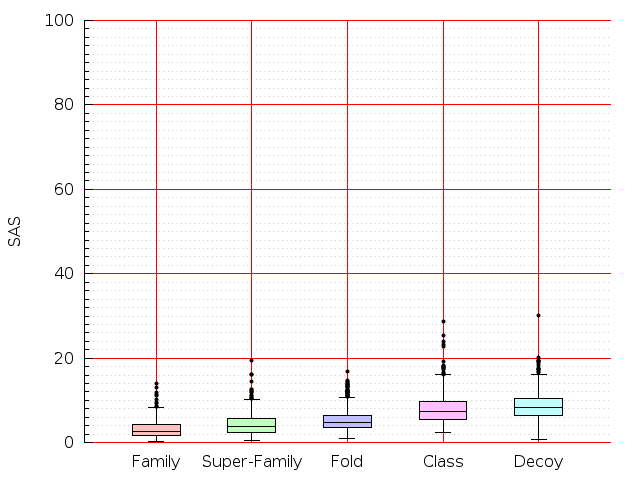 | 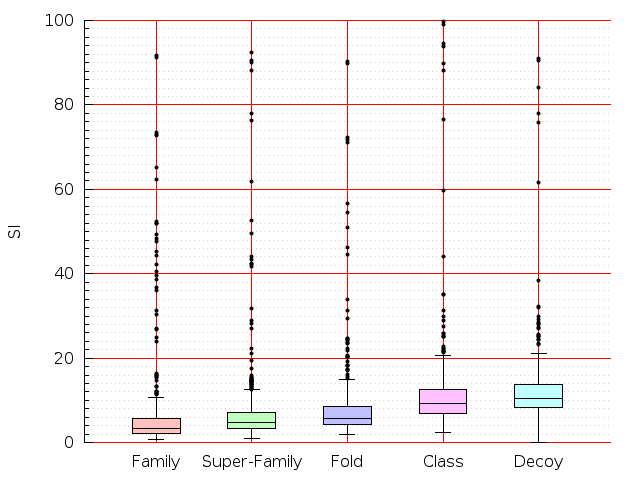 | 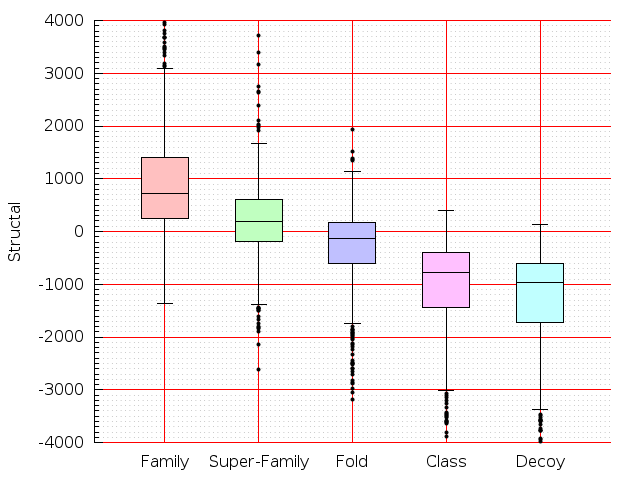 | 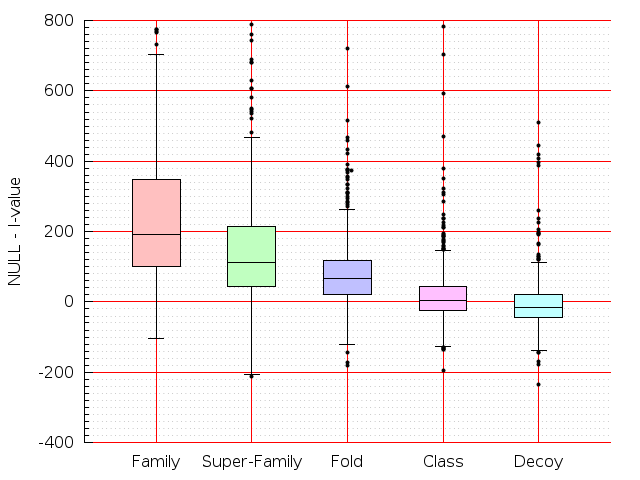 | TM-Align |
| CE | 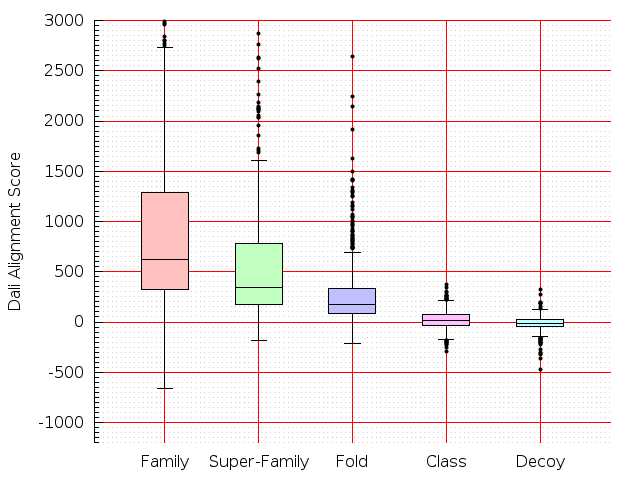 | 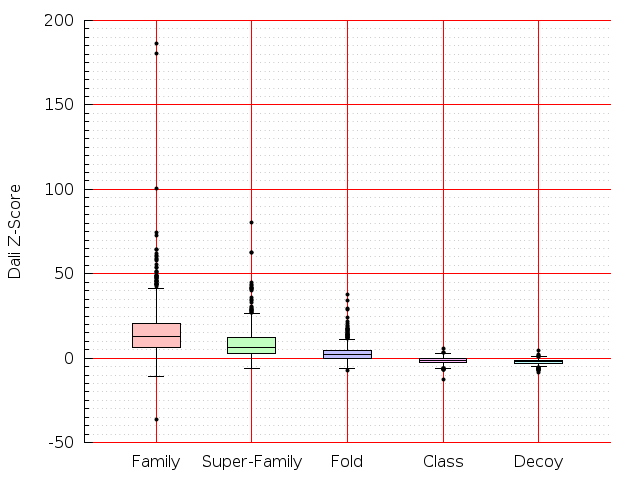 | 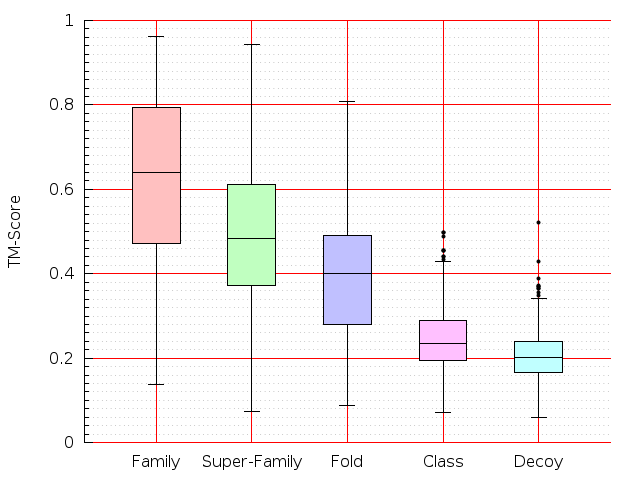 | 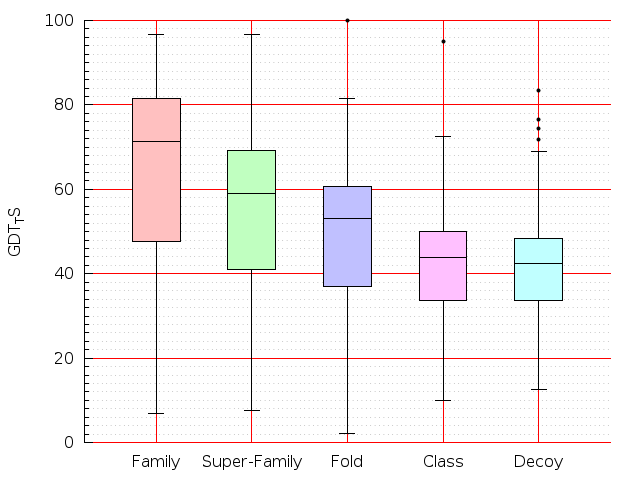 | 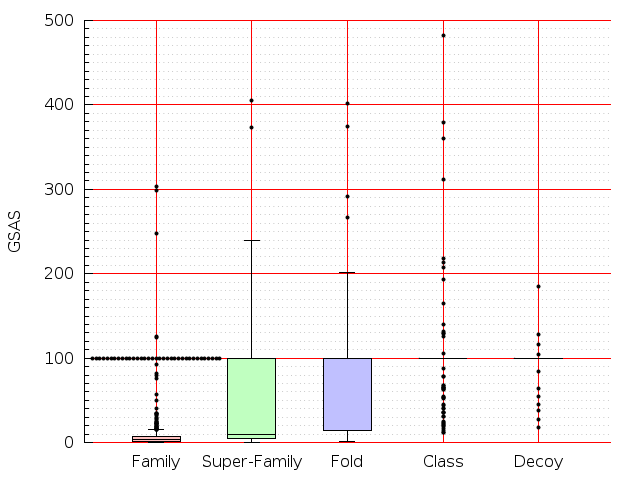 | 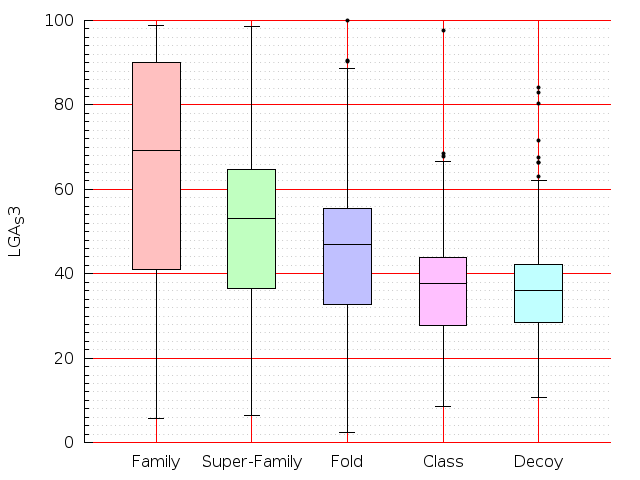 | 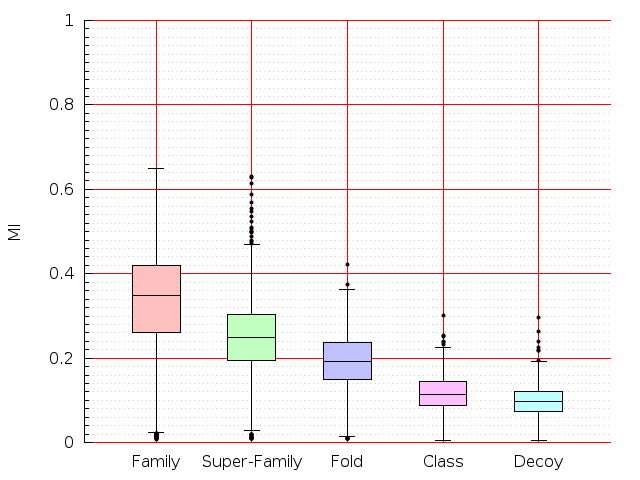 | 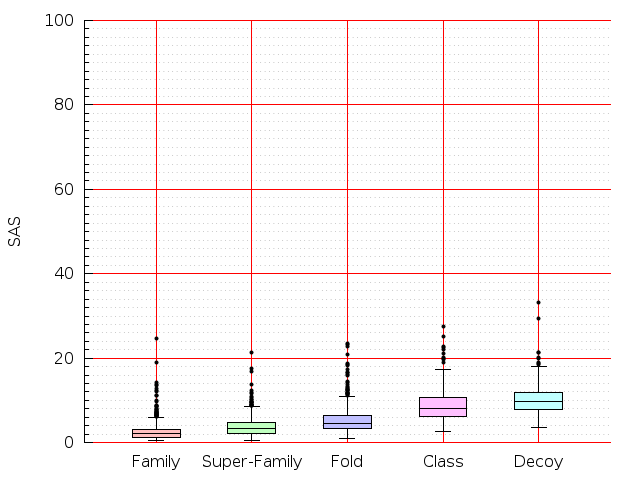 | 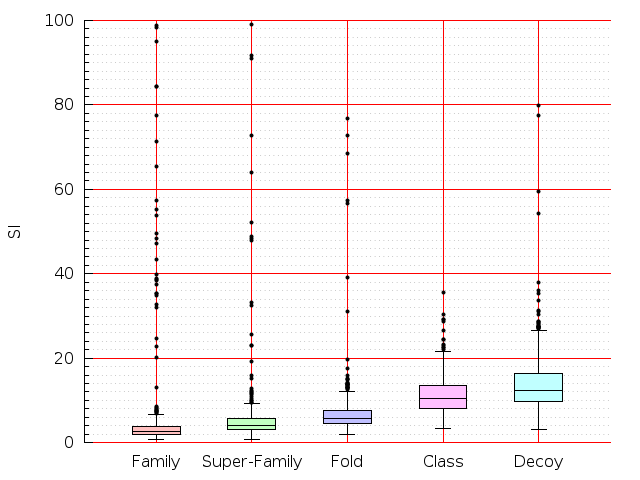 | 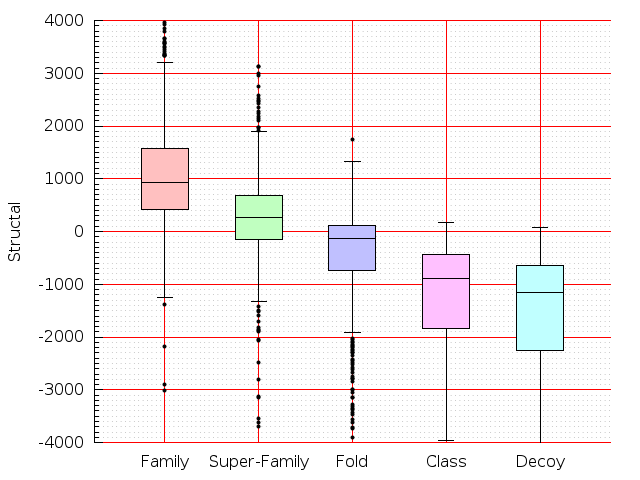 | 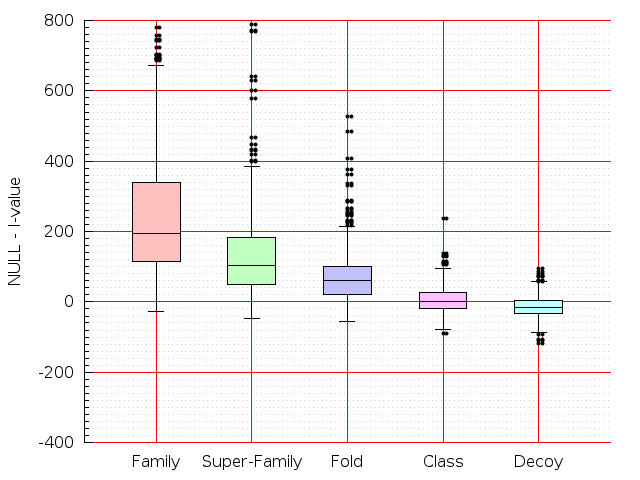 | CE |
| FatCat | 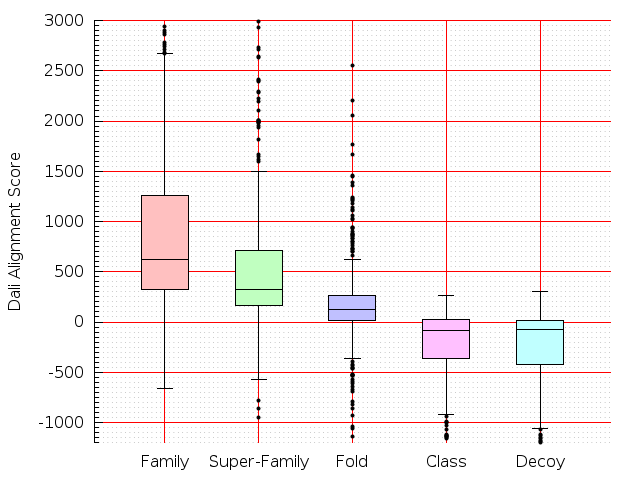 | 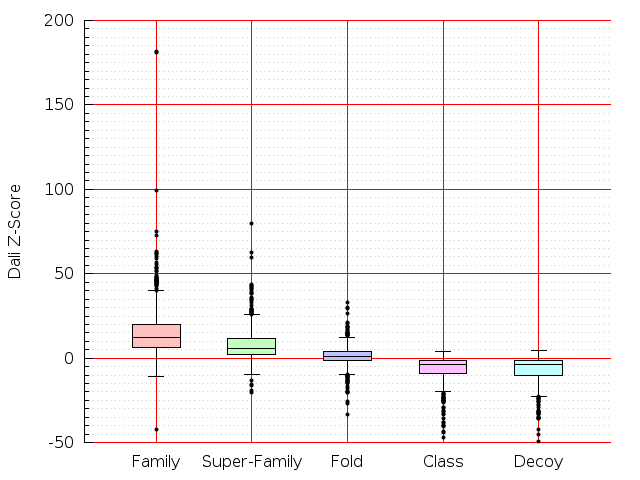 | 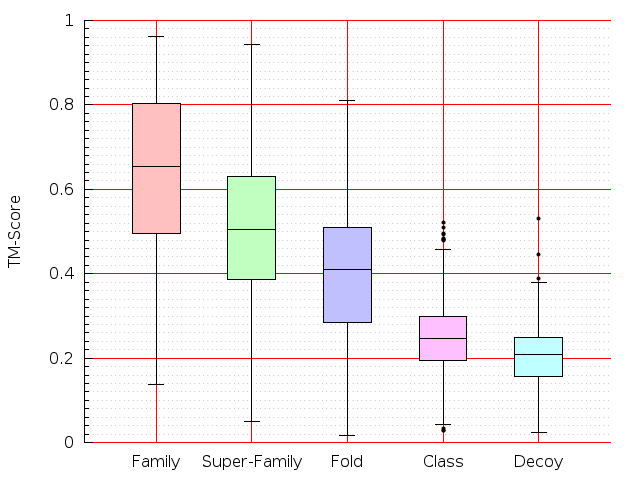 | 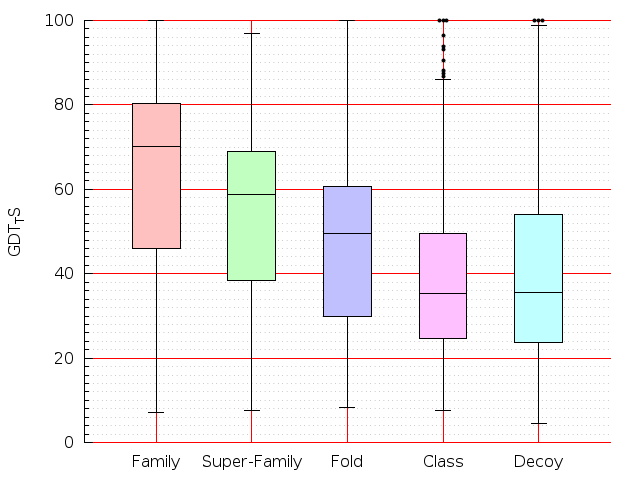 | 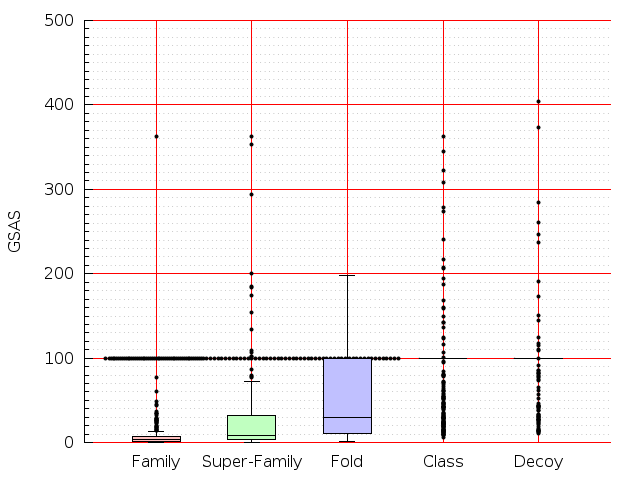 | 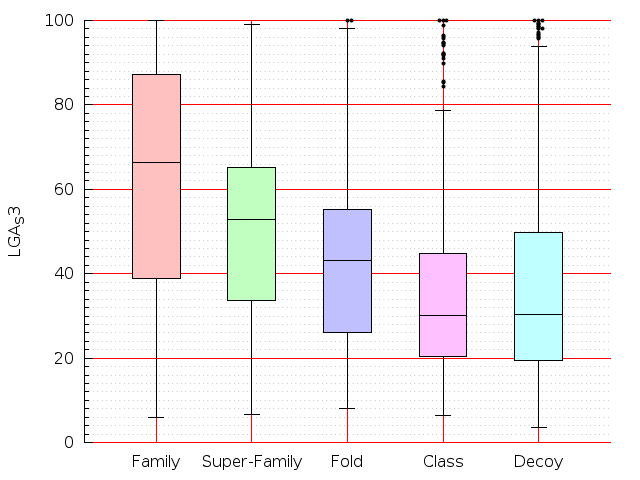 | 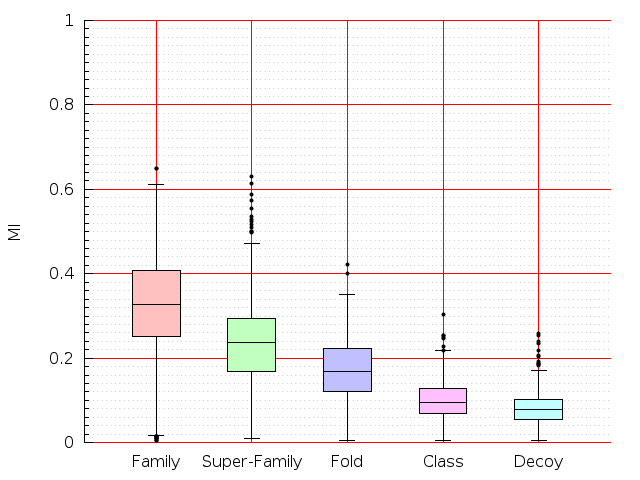 | 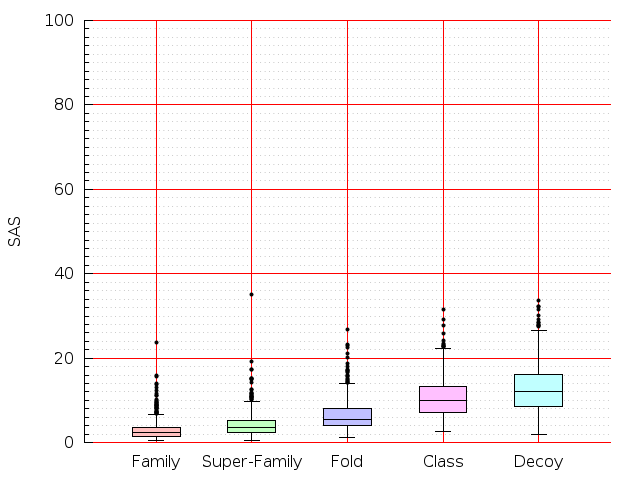 | 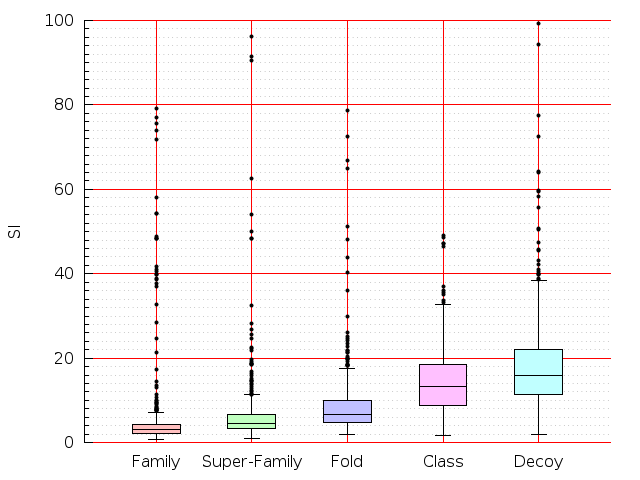 | 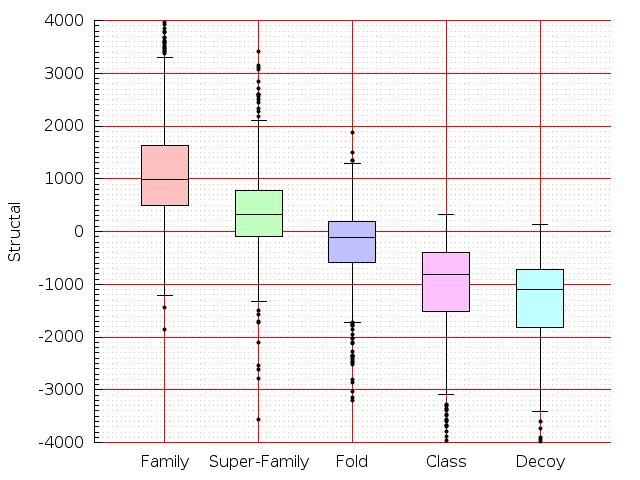 | 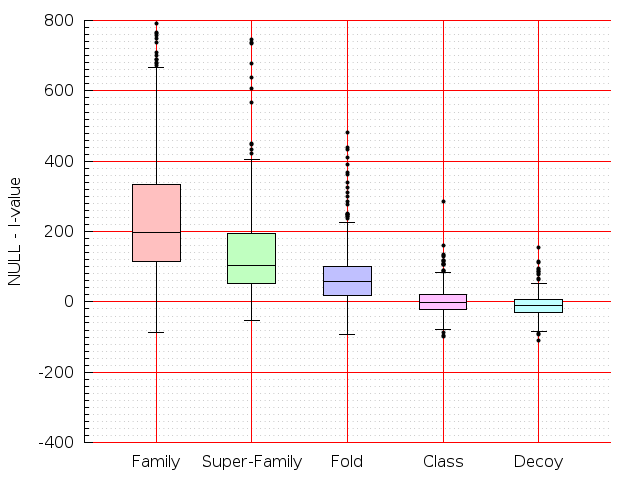 | FatCat |
| LGA | 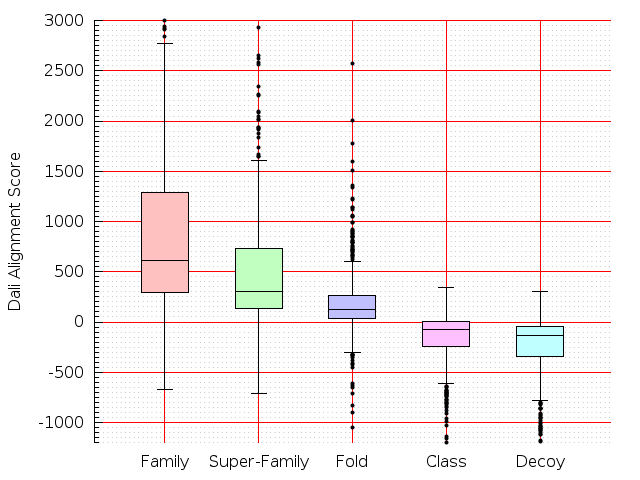 | 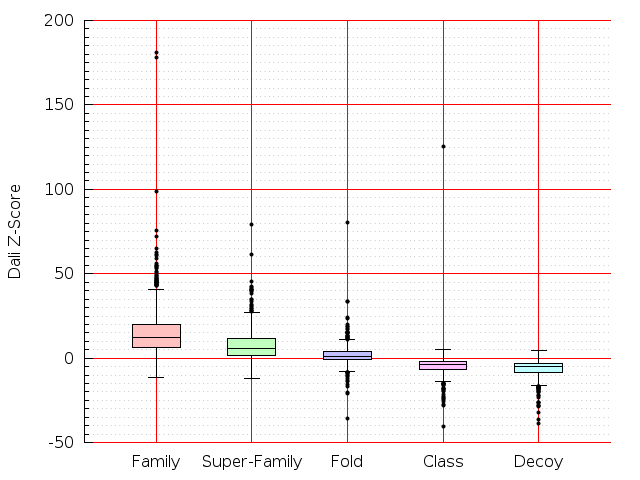 | 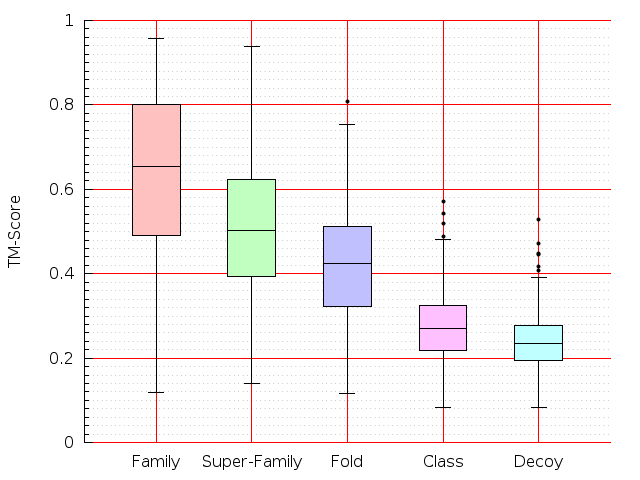 | 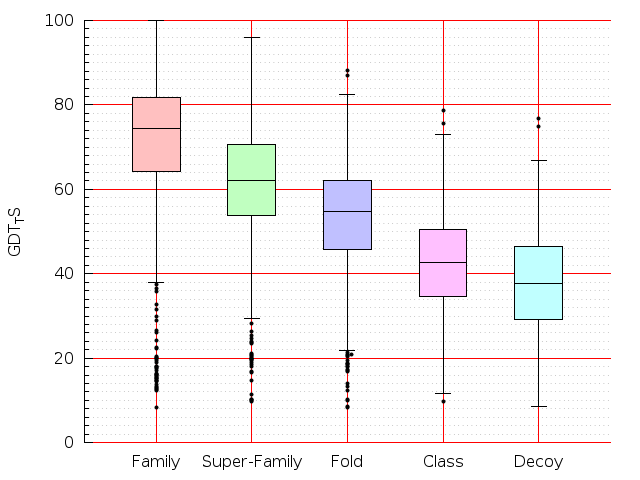 | 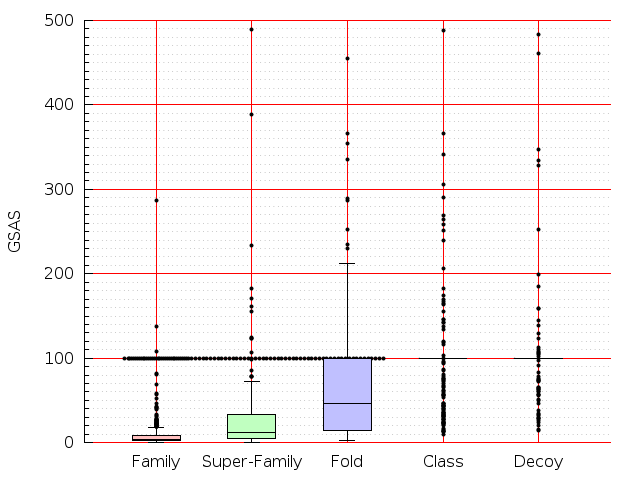 | 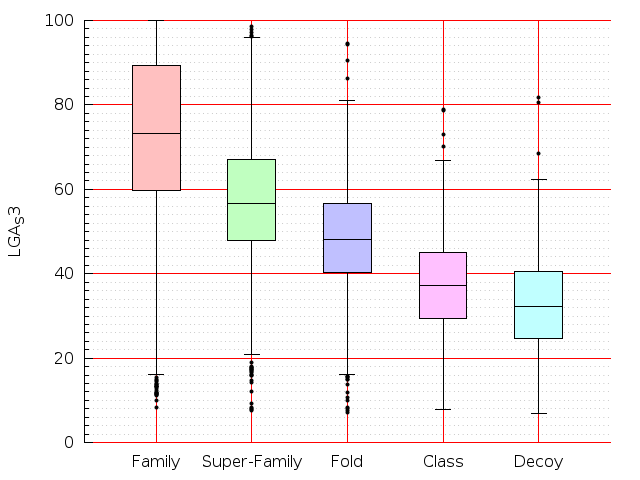 | 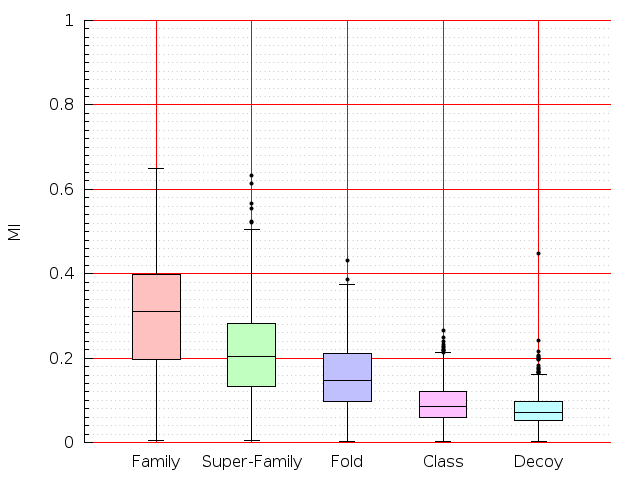 | 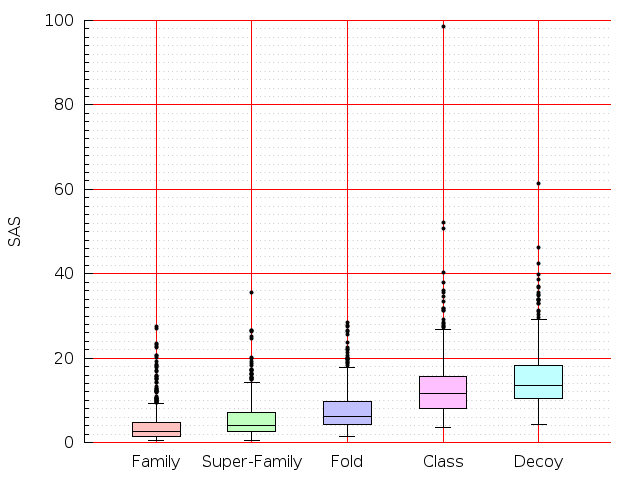 | 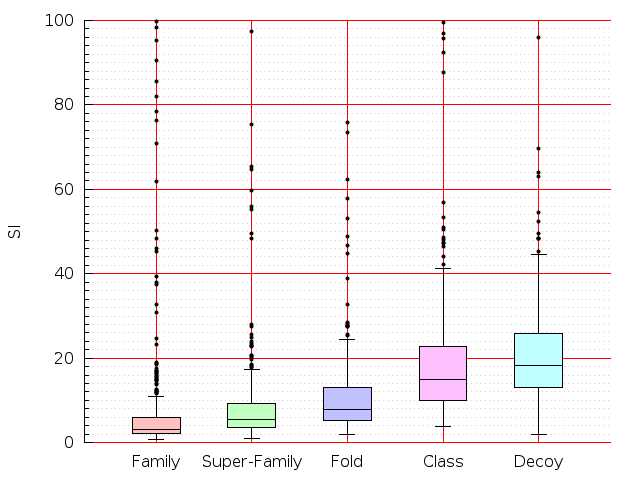 | 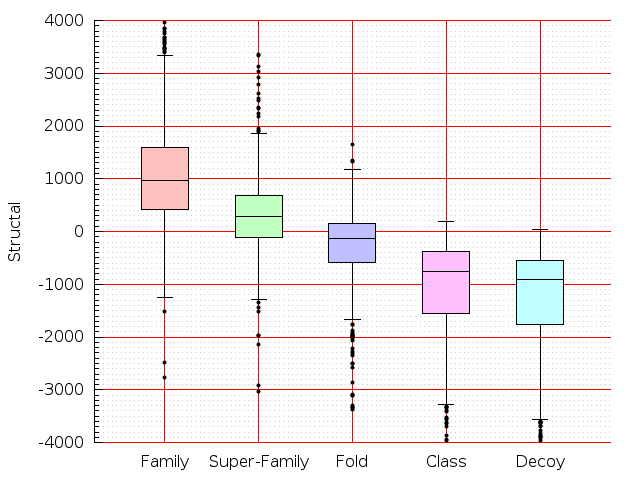 | 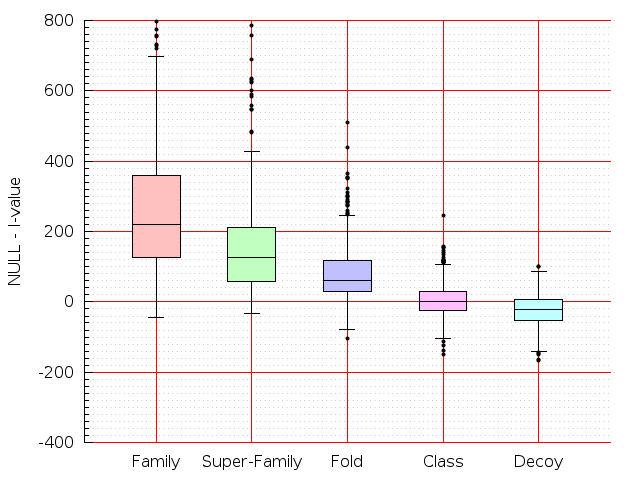 | LGA |
| Dali Score | TM-Score | GDT_TS | GSAS | LGA_S3 | MI | SAS | SI | Structal Score | I-value |
Table S2: Listing of disagreeing pairs
This table contains a full listing off all disagreeing pairs of scores. You may find that some files are empty, this occurs when the pair of scores agree on the quality ranking across all aligned pairs. Clicking on a shaded cell in the table allows you to download the listing of disagreement between two scoring functions indicated by the column/row headings.
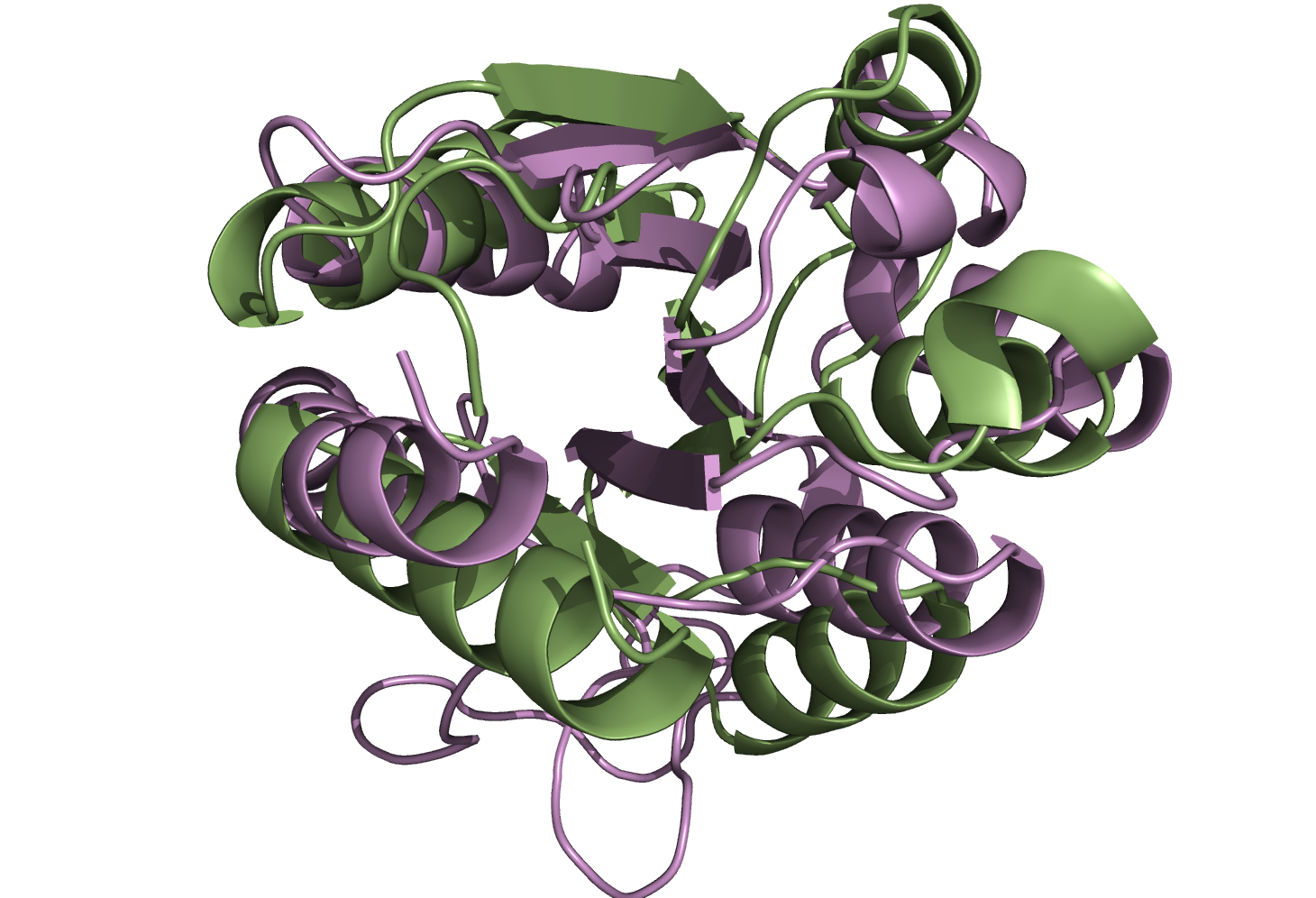
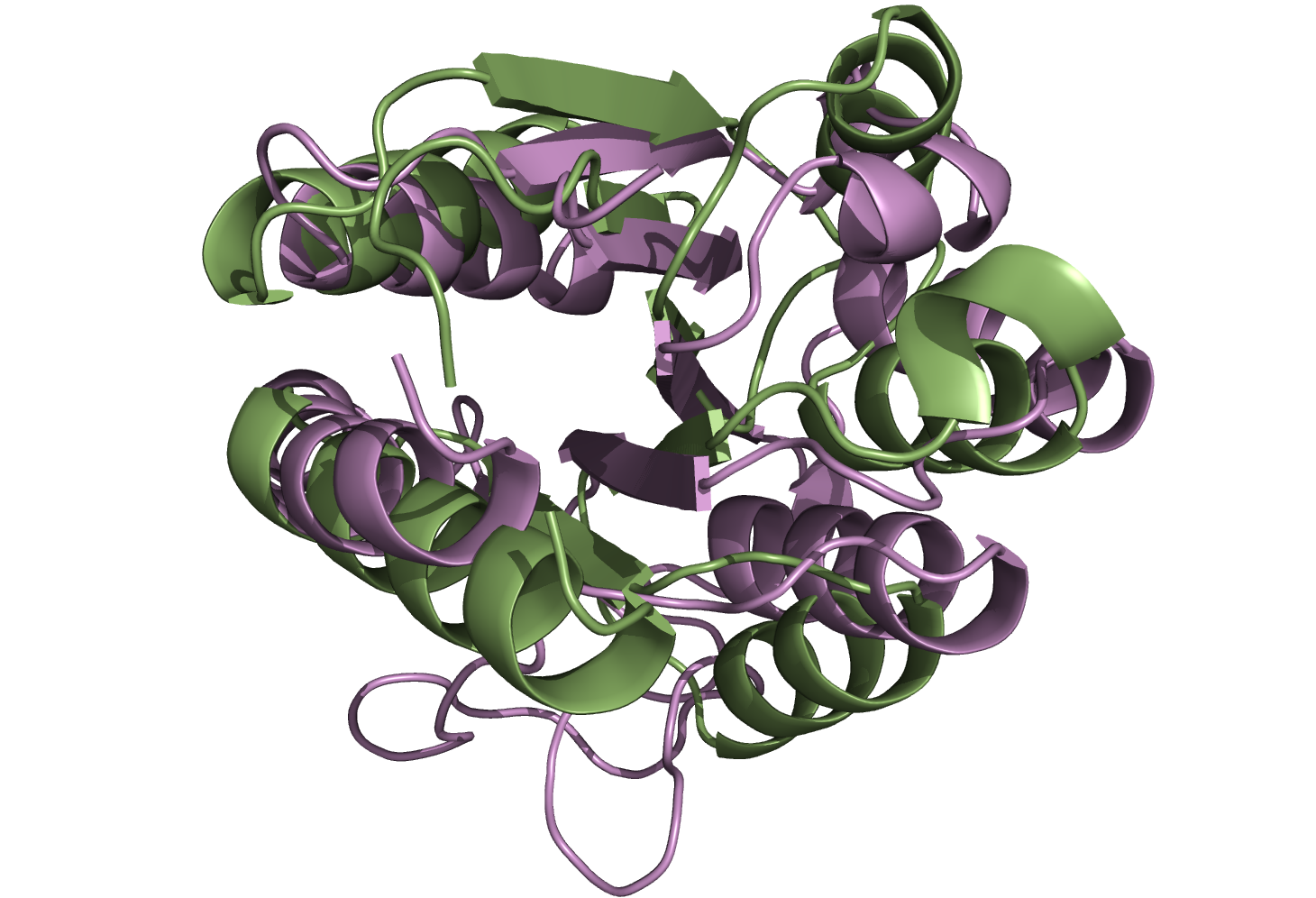
Superpositions from Figure 3 in the main text
Superpositions of PDB:1RCF and PDB:3CHY selected as the best by DALI (left) and I-value (right) from sieved alignments.